Automated Image-Based Quantification of Neutrophil Extracellular Traps Using NETQUANT | Protocol (Translated to Hebrew)
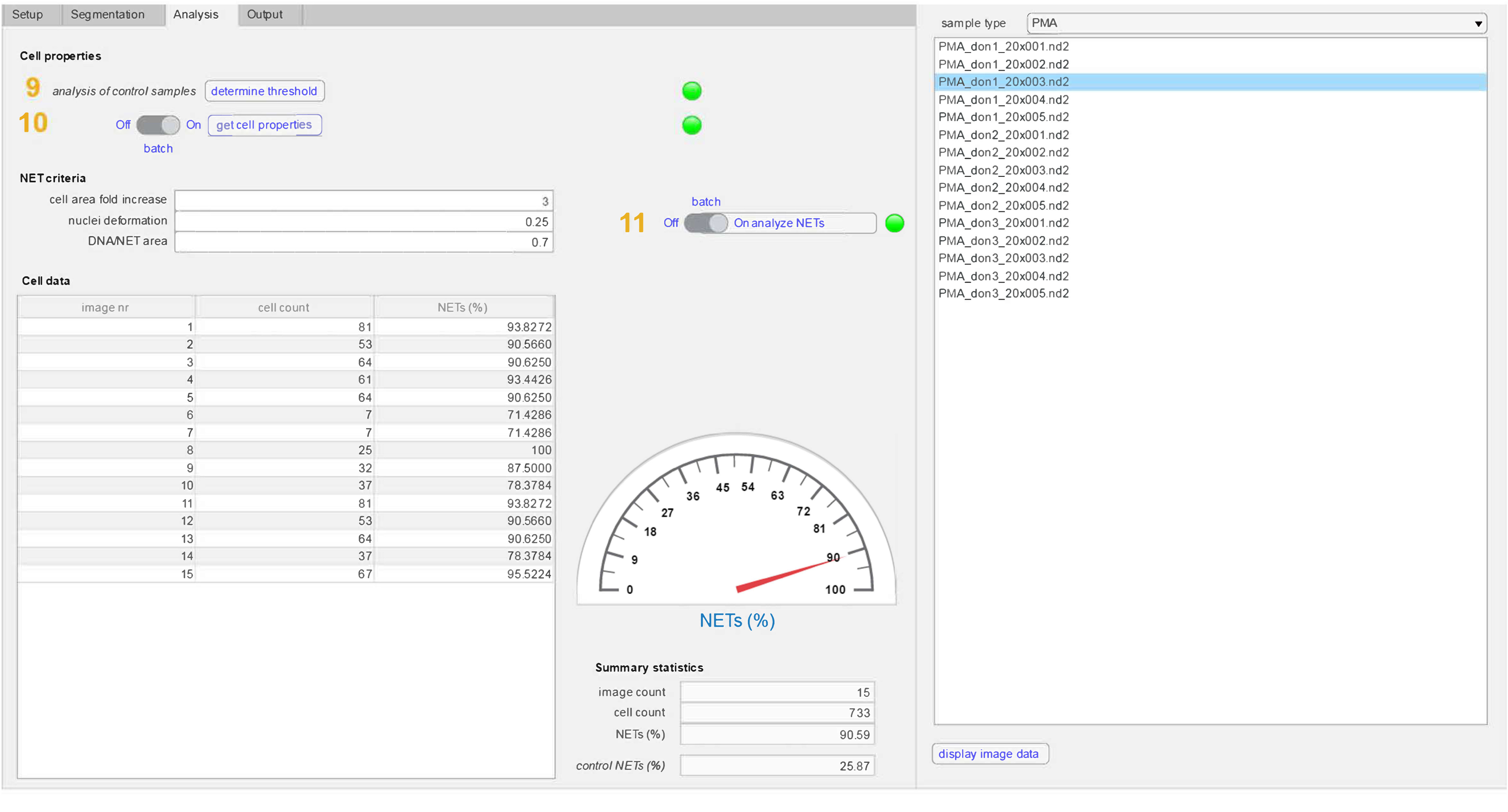
Automated Image-Based Quantification of Neutrophil Extracellular Traps Using NETQUANT | Protocol (Translated to Hebrew)
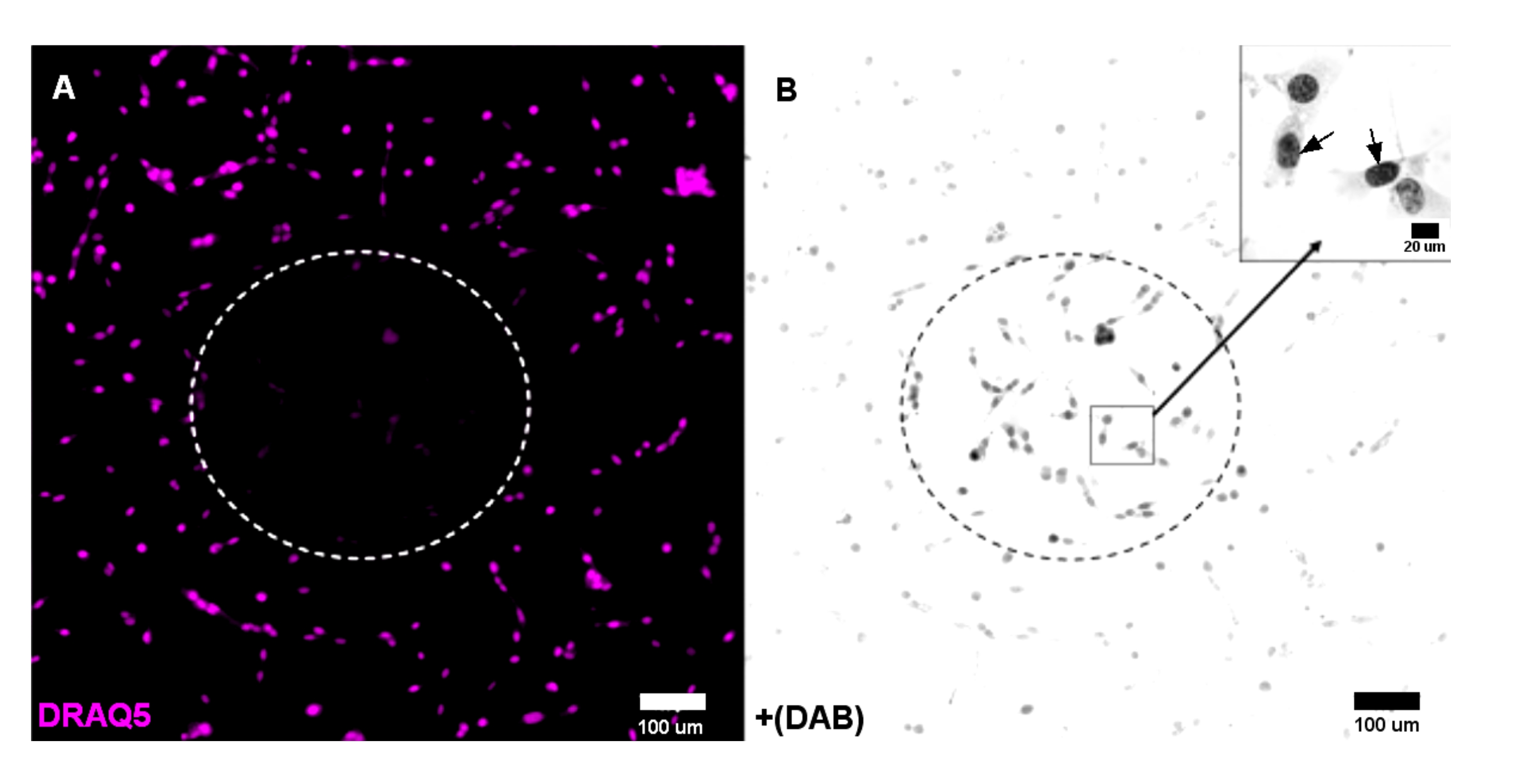
Imaging Replicative Domains in Ultrastructurally Preserved Chromatin by Electron Tomography | Protocol (Translated to Hebrew)
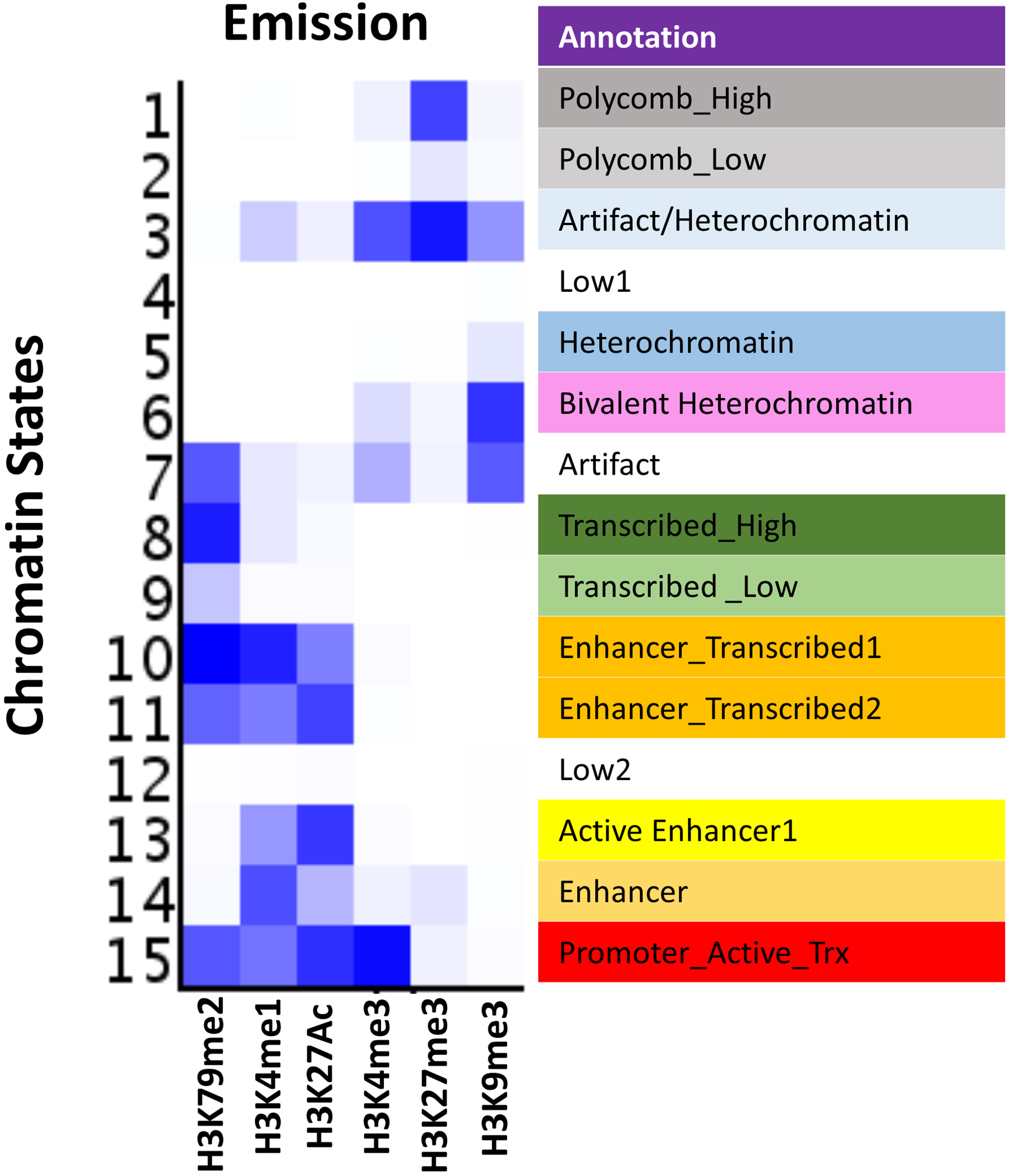
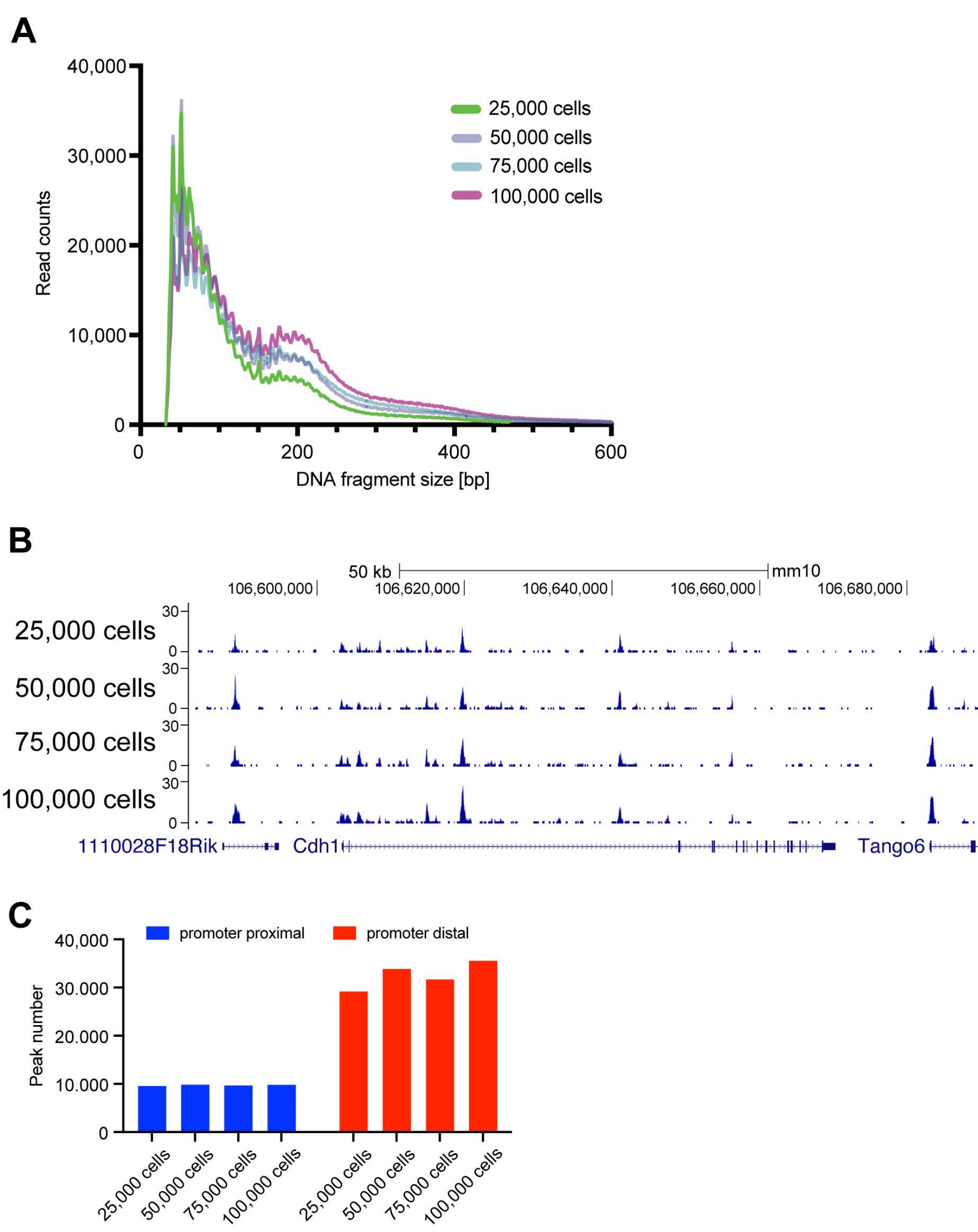
An Integrated Platform for Genome-wide Mapping of Chromatin States Using High-throughput ChIP-sequencing in Tumor Tissues | Protocol (Translated to Hebrew)

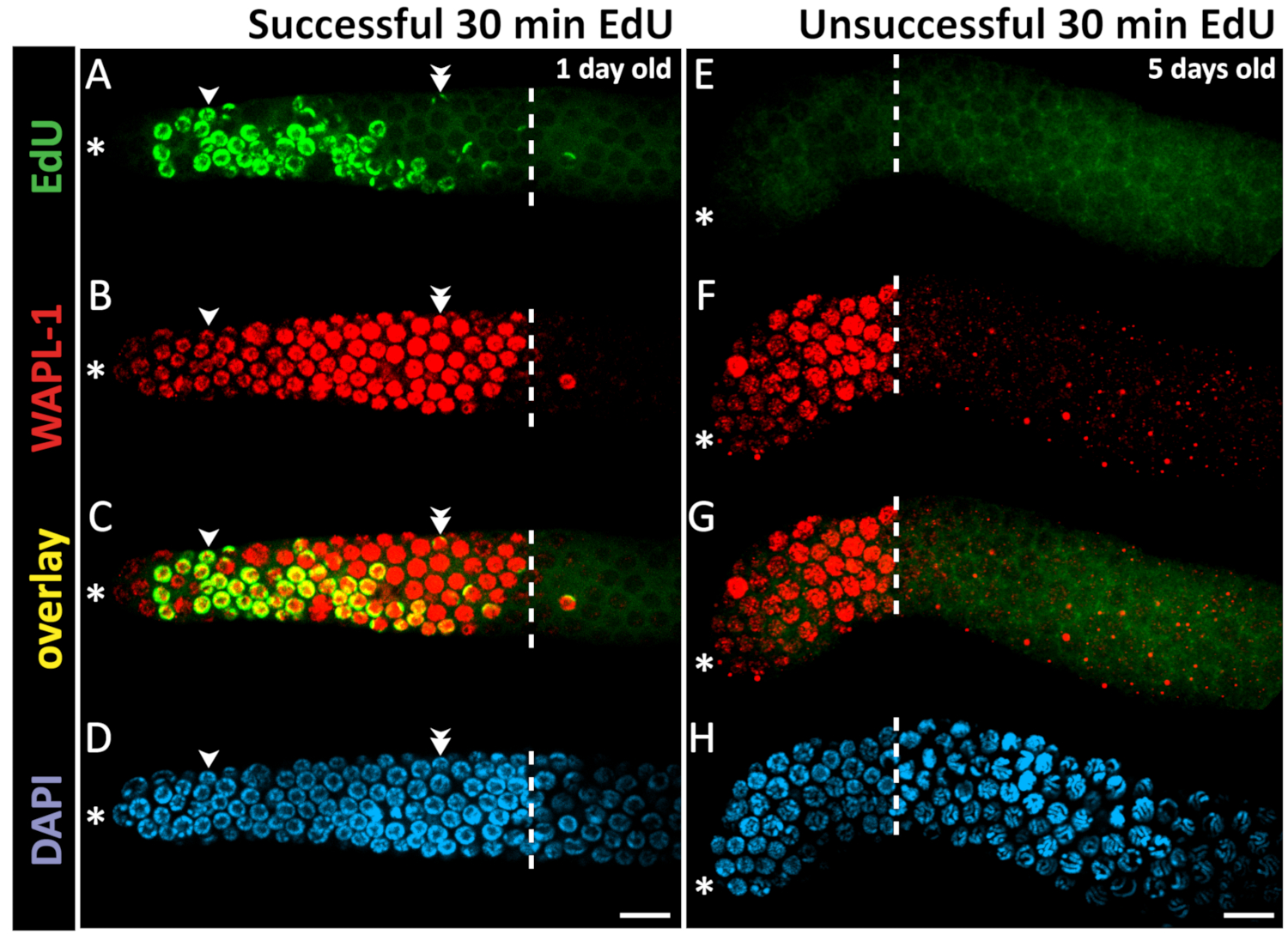
Chromatin Spread Preparations for the Analysis of Mouse Oocyte Progression from Prophase to Metaphase II | Protocol (Translated to Hebrew)

Automated Image-Based Quantification of Neutrophil Extracellular Traps Using NETQUANT | Protocol (Translated to Hebrew)

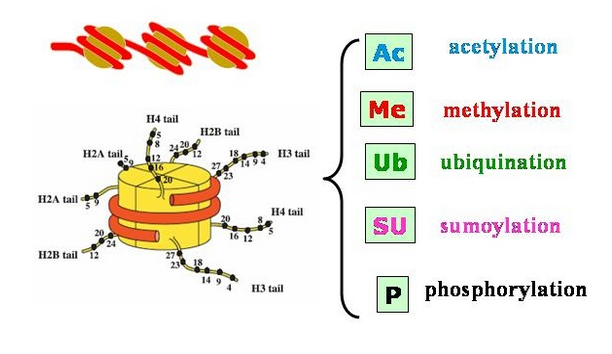
Chromatin Immunoprecipitation (ChIP) of Histone Modifications from Saccharomyces cerevisiae | Protocol (Translated to Hebrew)
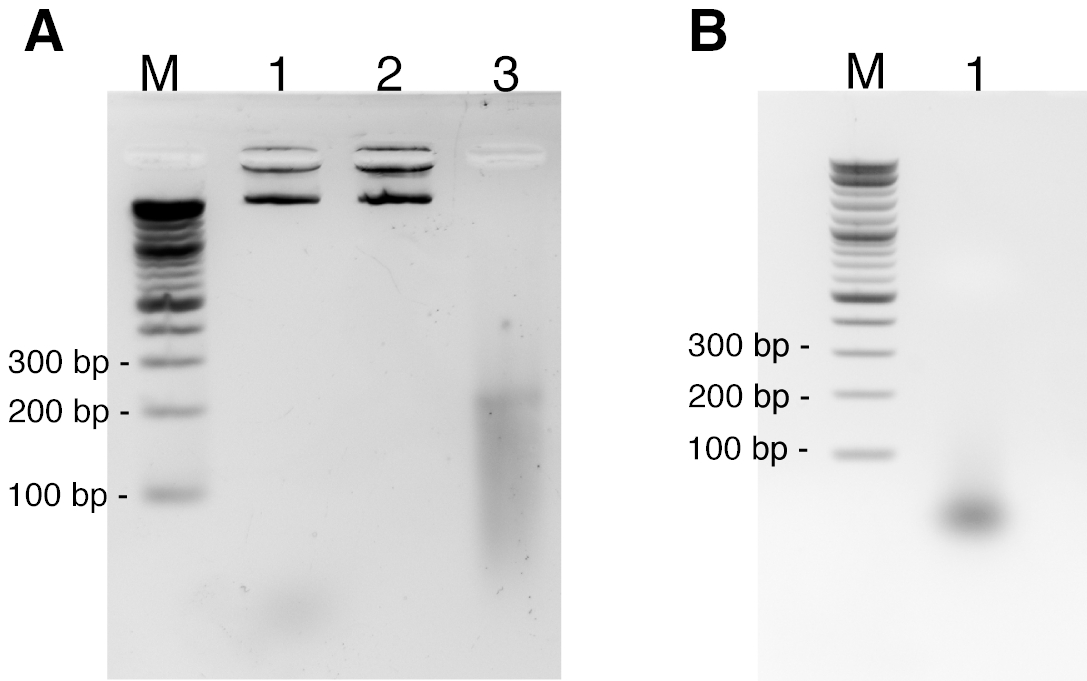
3D Multicolor DNA FISH Tool to Study Nuclear Architecture in Human Primary Cells | Protocol (Translated to Hebrew)
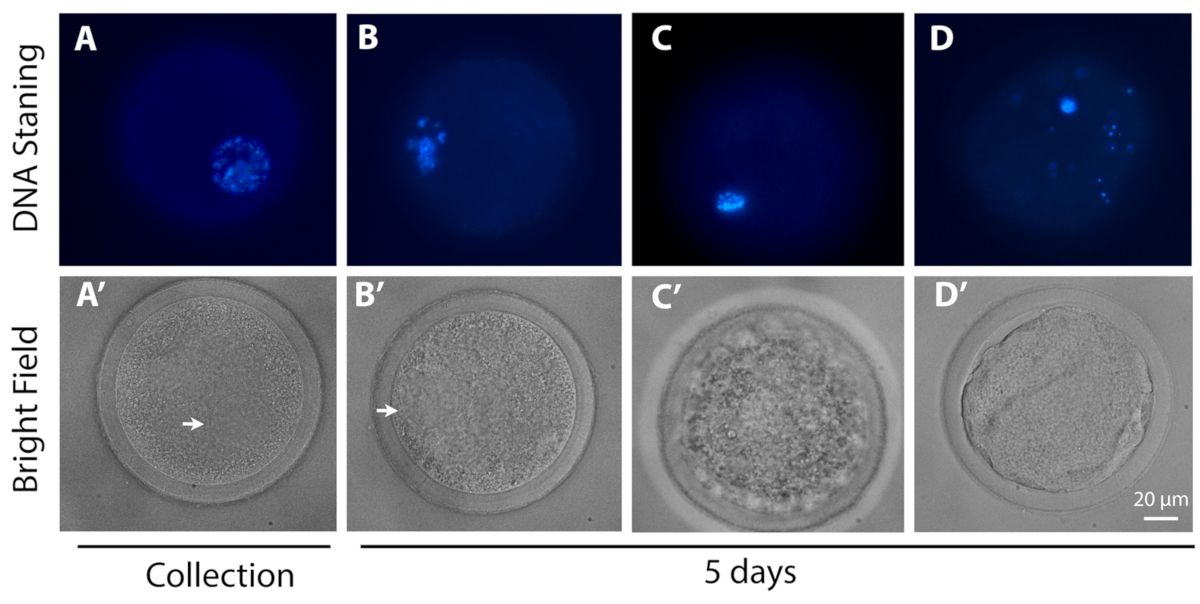
In Vitro Culture Strategy for Oocytes from Early Antral Follicle in Cattle | Protocol (Translated to Hebrew)
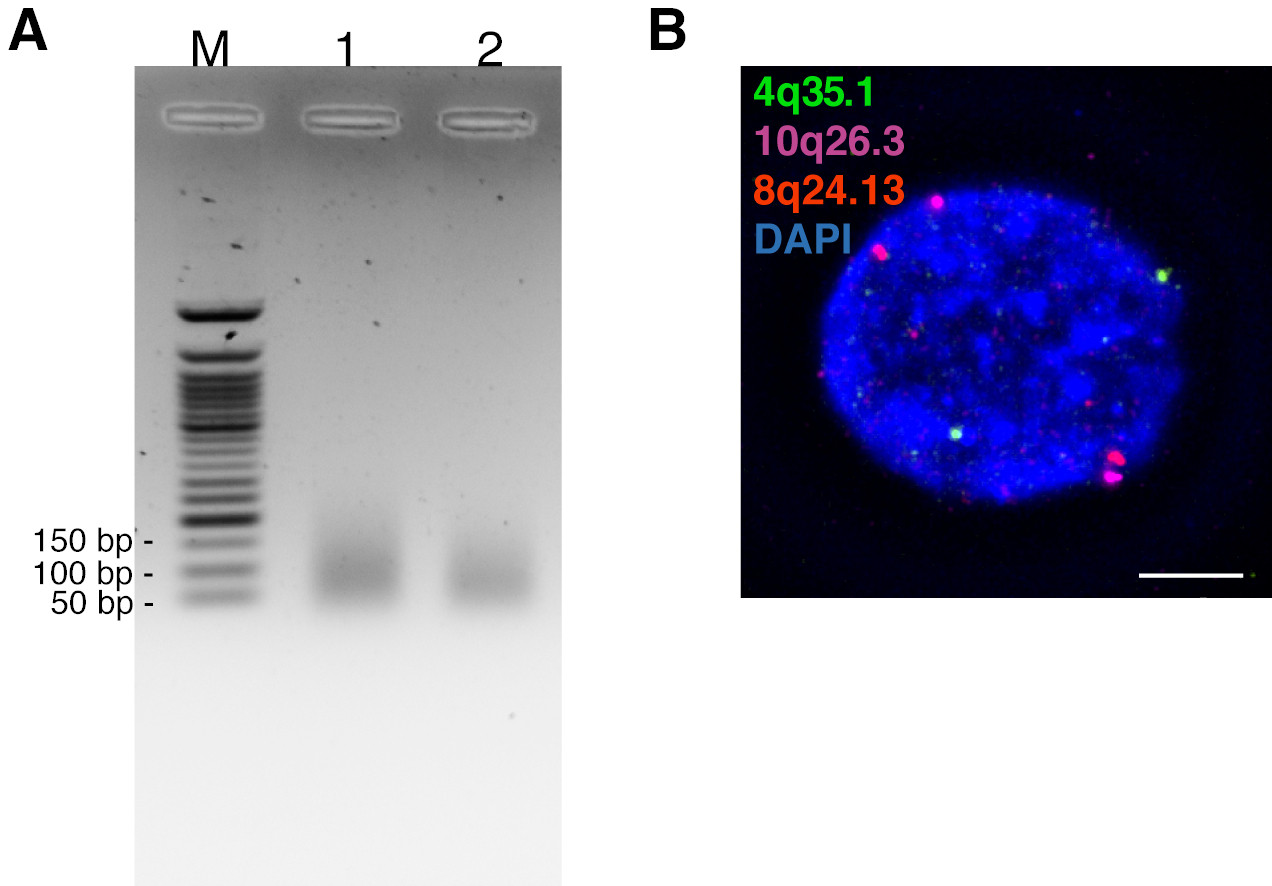
3D Multicolor DNA FISH Tool to Study Nuclear Architecture in Human Primary Cells | Protocol (Translated to Hebrew)
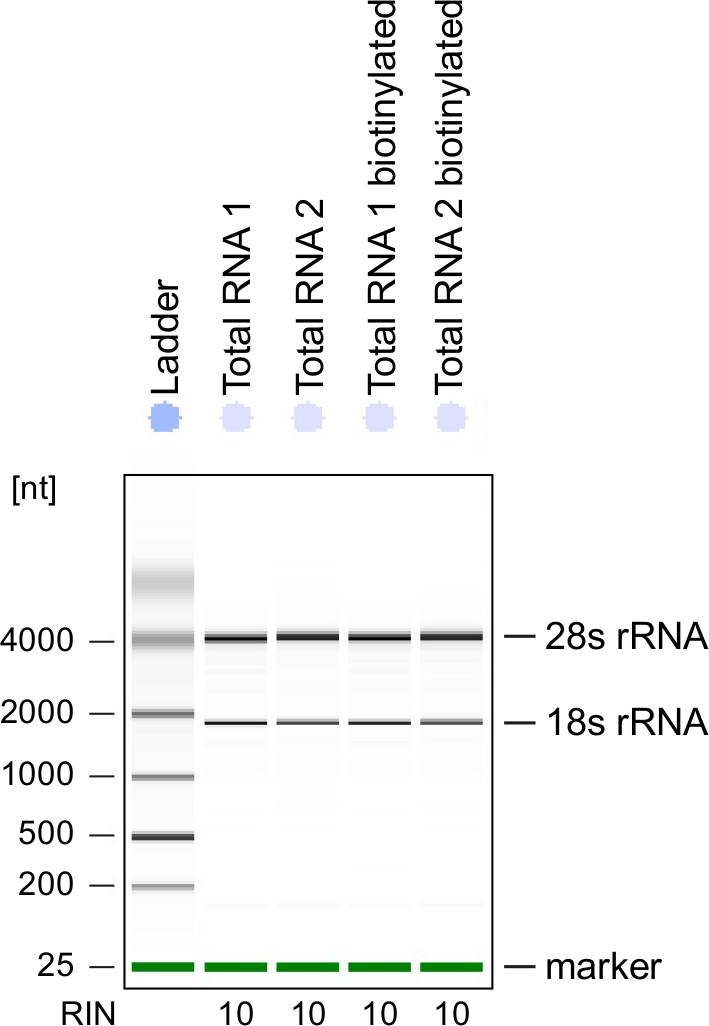
Real-time Analysis of Transcription Factor Binding, Transcription, Translation, and Turnover to Display Global Events During Cellular Activation | Protocol (Translated to Hebrew)